필요 데이터
- phyloseq
목표 결과
- 각 alpha diversity 간의 통계 분석 자료
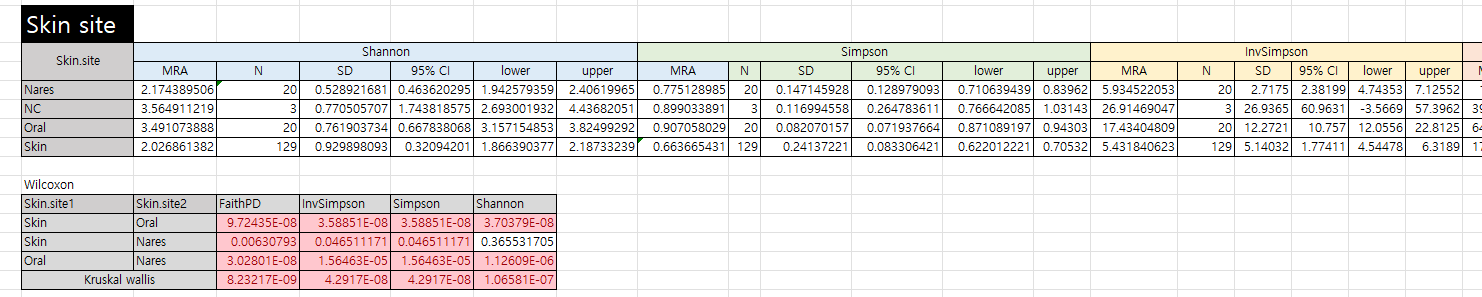
제작 이유
- 엑셀로 정리하다가 코딩으로 만들면 훨씬 간편할 것 같아서
- 다 만들고 나니 정말로 그랬다.
#### 필요
library(phyloseq)
library(dplyr)
library(picante)
#### import data
ps = phyloseq
#### make diversity table
Alpha.Shannon <- estimate_richness(ps, measures = "Shannon")
Alpha.Simson <- estimate_richness(ps, measures = "Simpson")
Alpha.InvSimson <- estimate_richness(ps, measures = "InvSimpson")
Alpha.FaithPD = t(picante::pd(samp = otu_table(ps), tree = phy_tree(ps), include.root = F))[1,]
Alpha.FaithPD.2 <- data.frame(`FaithPD` = Alpha.FaithPD)
Alpha.1 <- merge(Alpha.Shannon, data.frame(sample_data(ps)) , by = "row.names") %>%
column_to_rownames("Row.names")
Alpha.2 <- merge(Alpha.Simson, Alpha.1, by = "row.names") %>%
column_to_rownames("Row.names")
Alpha.3 <- merge(Alpha.InvSimson, Alpha.2, by = "row.names") %>%
column_to_rownames("Row.names")
Alpha.4 <- merge(Alpha.FaithPD.2, Alpha.3, by = "row.names") %>%
column_to_rownames("Row.names")
# 사실 cbind해도 되는데, 혹시 순서 바뀔까봐 merge로 바꿈
#### save total diversity
write.csv(Alpha.4, paste0(path, "alpha_data.csv"), col.names = T, row.names = T)
#### function
alpha_df <- function(df = df,
alpha_aiversity = c("FaithPD", "InvSimpson", "Simpson", "Shannon"),
group = group,
path = path) {
# setting
options(scipen = 999) # 숫자 요약에서 풀로 보기
compar <- as.vector(unique(df[, group])) # 비교하고자 하는 그룹
compar_num <- length(compar) # 비교하고자 하는 그룹 수
permut <- combinat::combn(compar, 2) %>% t() # 순열
p_val_df <- permut %>% as.data.frame() # p-value값이 들어갈 data.frame
num <- nrow(permut)
range <- seq(from = 1, to = num, by = 1) # for문을 위한 수열
for (index in alpha_aiversity) { # 각 alpha diversity index동안
# normal distribution : https://stackoverflow.com/questions/35953394/calculating-length-of-95-ci-using-dplyr
result <- df %>%
dplyr::group_by(!!rlang::sym(group)) %>%
dplyr::summarize(
Mean = mean(!!rlang::sym(index)), # 평균
N = n(), # 행 개수
Sd = sd(!!rlang::sym(index))) %>% # 표준편차
dplyr::mutate(se = Sd /sqrt(N),
lower = Mean - qnorm(0.975)*se, # 95% 신뢰 구간 하한
upper = Mean + qnorm(0.975)*se, # 95% 신뢰 구간 상한
CI95per = upper-lower # 95% 신뢰 구간
)
assign(paste0(index, "_df"), result)
if (compar_num>=3){ # 3군 이상일때 kruskal wallis 분석
### kruskal wallis
krus_p <- kruskal.test(df[, index] ~ df[, group], data = df)$p.value
p_val_df["kruskal wallis—pval", index] <- krus_p
}
for (rows in range) { # 2 군씩 나누어서 pairwise wilcoxon rank sun test
### wilcoxon
compar_var <- permut[rows, ]
compar_df <- df %>% filter((!!rlang::sym(group)) %in% compar_var)
p_val_df[rows, index] <- wilcox.test(compar_df[, index] ~ compar_df[, group], data = compar_df)$p.value
}
p_val_df[, paste0(index, "_adjust")] <- p.adjust(p_val_df[, index], method = "fdr")
}
#### Save_mean value
Total_result <- data.frame(
Shannon = Shannon_df,
Simpson = Simpson_df,
InvSimpson = InvSimpson_df,
FaithPD = FaithPD_df )
write.csv(Total_result, paste0(path, "alpha_",group , "_mean_and_CI.csv"), col.names = T, row.names = T)
#### Save_statistic value
print(p_val_df)
write.csv(p_val_df, paste0(path, "alpha_",group , "_StatisticTest.csv"), col.names = T, row.names = T)
}
밤 샘의 결과물..
반응형
